|
Multi-factorial statistics control for batch effects and support paired studies. All tools account for differences due to sequencing depth, removing the need to normalize input data. The free “Advanced RNA-Seq plugin” integrates all the analysis steps – from secondary analysis of the reads to sophisticated statistics – into easy-to-use workflows, and gives access to a wide range of experimental designs, from case-control or multi-group experiments to multi-factorial experiments. If assembly of long reads (like PacBio) or genome finishing are the primary focus, then CLC Genomics Workbench can be enhanced with the commercial CLC Genome Finishing Module. Like for our read mapper, a wide range of NGS data types is supported, and hybrid assemblies combine the unique strength of short and long reads for optimal results. Our trusted de novo assembler accompanied by trimming tools to remove low quality data deliver assembly quality fast and compute resource efficient. All made accessible through our tried and trusted interactive graphical user interface. Transcription factor ChIP-seq exposes defined peak regions characteristic of transcription binding sites throughout the whole genome. Recent improvements promise deeper insights into transcriptional regulation even in the absence of control sample. Our goal is to reduce your manual work and focus on deriving biological meaningful results from raw NGS reads. Local realignment can drastically reduce false positive detection rates for certain variant types. It also supports the use of hybrid data sets. The algorithm offers comprehensive support for a variety of data formats, including both short and long reads, and all flavors of paired read data regardless of insert size or read orientation. 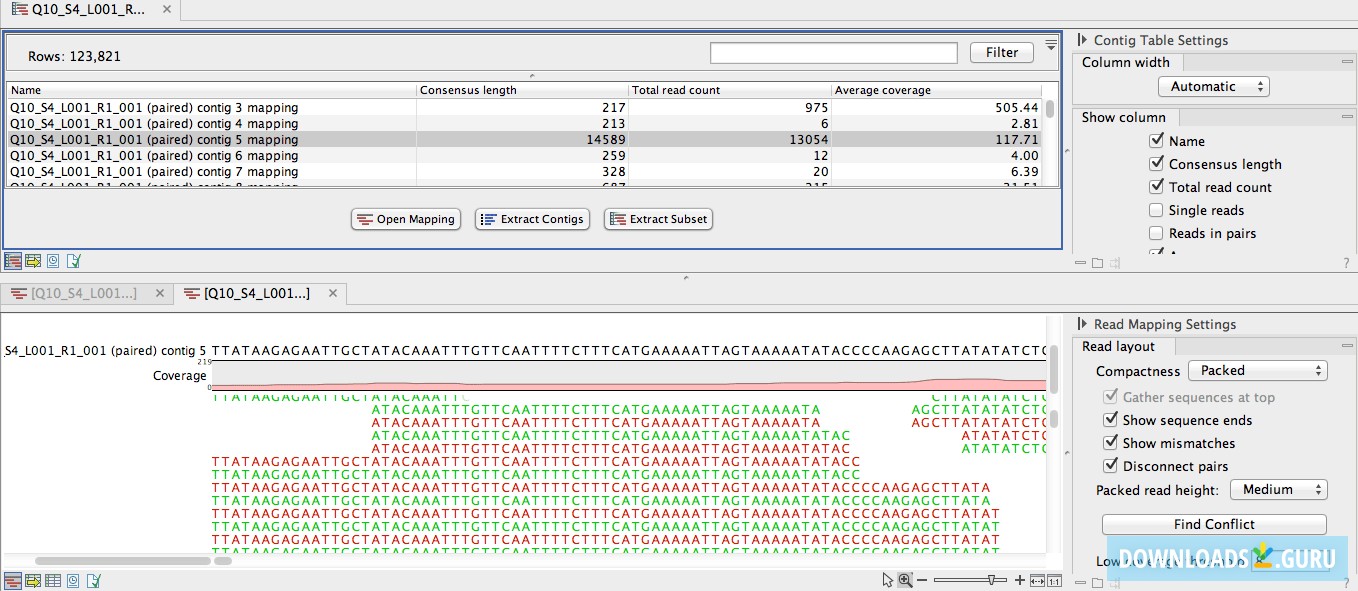
Our algorithm is optimized for high-quality mapping of large data volumes in a fast and memory-efficient way. The first step in resequencing is accurate read mapping. When dealing with high sample volumes efficient algorithms reduce run time while customizable analysis workflows and batch processing shorten hands-on time to a minimum.ĬLC Genomics Workbench allows you to focus on the biological interpretation of detected variants. Its cutting-edge technology incorporates unique features and algorithms that are widely used by scientific leaders in industry and academia to overcome bottleneck challenges associated with data analysis.ĬLC Genomics Workbench supports the complete resequencing pipeline for detecting and comparing genetic variants. No license required!ĬLC Genomics Workbench is a powerful solution developed by scientists for scientists to analyze and visualize next generation sequencing (NGS) data. 
BMC Genomics 15:15īolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data.With version 11, CLC Genomics Workbench can now be used as a free genome browser to share, view, and explore NGS analysis results.Įnjoy support for a wide range of open and proprietary file formats. Hou CY, Wu MT, Lu SH, Hsing YI, Chen HM (2014) Beyond cleaved small RNA targets: unraveling the complexity of plant RNA degradome data. Lin PC, Lu CW, Shen BN, Lee GZ, Bowman JL, Arteaga-Vazquez MA, Liu LY, Hong SF, Lo CF, Su GM, Kohchi T, Ishizaki K, Zachgo S, Althoff F, Takenaka M, Yamato KT, Lin SS (2016) Identification of miRNAs and their targets in the liverwort Marchantia polymorpha by integrating RNA-Seq and degradome analyses. Zhai J, Arikit S, Simon SA, Kingham BF, Meyers BC (2014) Rapid construction of parallel analysis of RNA end (PARE) libraries for Illumina sequencing. Niu QW, Lin SS, Reyes JL, Chen KC, Wu HW, Yeh SD, Chua NH (2006) Expression of artificial microRNAs in transgenic Arabidopsis thaliana confers virus resistance. Souret FF, Kastenmayer JP, Green PJ (2004) AtXRN4 degrades mRNA in Arabidopsis and its substrates include selected miRNA targets.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
